Supplementary Figure 1. Representative controls for DSB induction efficiency in wild-type
|
|
|
- Leslie Fitzgerald
- 5 years ago
- Views:
Transcription
1 Supplementary Figure 1. Representative controls for DSB induction efficiency in wild-type and mutant cells. Representative spotting assay reveals that the tested deletions do t affect the efficiency of DSB induction, which results in the loss of the URA3 marker gene. Shown are ten-fold serial dilutions of cells with indicated getypes onto n-selective media (SC), negative selection media (SC-URA; DSB induction indicates less growth), and positive selection media (SC+5FOA; DSB induction indicates more growth). DSB efficiency is consistently above 99.9% in all strains tested. S1
2 Supplementary Figure 2. ChIP assessing chromatin silencing near the subtelomeric DSB site. Representative anti-diach3 (K9, K14) ChIP gels revealing that the deletion of Lrs4, but t Cik1 or Nup84, greatly compromises silent chromatin assembly near the uninduced DSB site. Relative fold enrichments for regions surrounding the DSB site are presented below the gel. Results are rmalized to ACT1 and presented relative to wild-type cells. S2
3 Supplementary Figure 3. DSB induction efficiency is similar between cells with or without zip/control sequences integrated near the subtelomeric DSB site. Representative spotting assays indicating that the tested zip/control deletions do t affect the efficiency of subtelomeric DSB induction in wild-type, lrs4, nup84, and cik1 cells. Shown are ten-fold serial dilutions of cells with indicated getypes onto n-selective media (SC) or negative selection media (SC- URA; DSB induction indicates less growth). DSB efficiency is consistently above 99.9% in all strains tested. S3
4 Supplementary Figure 4. ChIP assessing chromatin silencing near the subtelomeric DSB sites in the presence of zip or control sequences. Representative anti-diach3 (K9, K14) ChIP gels revealing that the deletion of Lrs4, but t Cik1 or Nup84, greatly compromises silent chromatin assembly near the uninduced DSB site. Relative fold enrichments for a region near the DSB site are presented below the gel. Results are rmalized to ACT1 and presented relative to wild-type control. S4
5 Supplementary Figure 5. Raw data for Western blots shown as cropped images in the main figures. Since the Western blots shown in the main figures are heavily cropped for presentation purposes as a space-saving measure, original larger images of developed films are shown here. The related main figure panel numbers are indicated above each film and the approximate cropped areas shown in the main figure panels are highlighted with dashed boxes. S5
6 Supplementary Table 1. Relative survival rates of TELXI-L subtelomeric DSB. Getype a,b Rate of survival c P value d Rate relative to WT WT (± 0.016) N/A 1.00 nup (± 0.004) < nup145c (± 0.005) < esc (± 0.010) < lrs (± 0.011) < csm (± 0.013) < lrs4 csm (± 0.017) < lrs4 esc (± 0.006) < nup84 esc (± 0.007) < nup84 lrs (± 0.008) < heh (± 0.024) nur (± 0.017) cik (± 0.013) < lrs4 cik (± 0.014) < kar (± 0.007) < lrs4 kar (± 0.003) < cin (± 0.005) < bub (± 0.013) rad (± 0.008) < lrs4 rad (± 0.002) < dnl (± 0.008) lrs4 dnl (± 0.008) < ku (± 0.035) pol (± 0.001) < dpb (± 0.010) sae (± 0.033) < sir (± 0.107) < sir (± 0.096) < sir3 rad (± 0.003) < slx (± 0.003) < slx (± 0.004) < rad (± 0.003) < swr (± 0.003) < cik1 swr (± 0.005) < tub (± 0.017) < cik1 tub (± 0.003) < a WT and/or cik1 rates are shown in Figs. 1b, 1f, 1g, and 4d for comparison. b Rates in Fig. 1c are repeated from Fig. 1b with superimposed colour-coded results of PCR survival analysis. c Standard deviations are shown between brackets. d P calculated using Student's t-test. S6
7 Supplementary Table 2. Relative survival rates of internal DSB on chromosome XI. Getype a Rate of survival b P value c Rate relative to WT WT (± ) N/A 1.00 rad (± ) ku (± ) < dnl (± ) < nup (± ) esc (± ) lrs (± ) csm (± ) heh (± ) nur (± ) cik (± ) sir (± ) < sir (± ) < slx (± ) < slx (± ) < a WT rates are shown in Figs. 1f and 1g for comparison. b Standard deviations are shown between brackets. c P calculated using Student's t-test. S7
8 Supplementary Table 3. List of strains used in this study. KMY# Original getype feature pkm97 KMY574 1 MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa KMY602 MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa + pkm97 [I-Sce I (gal induc), LEU2d, TRP1] KMY778 MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, nup84δ::kanr KMY827 MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, nup84δ::kanr KMY575 MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, nup145cδ::kanr KMY669 MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, nup145cδ::kanr KMY730 MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, esc1δ::hphr KMY756 MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, esc1δ::hphr KMY631 MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, lrs4δ::kanr KMY658 MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, lrs4δ::kanr KMY663 MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, csm1δ::kanr KMY678 MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, csm1δ::kanr KMY1583 KMY1663 KMY1142 KMY1152 KMY1145 KMY1196 KMY1121 KMY1139 MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, csm1δ::kanr, lrs4 ::HphR MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, csm1δ::kanr, lrs4 ::HphR MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, lrs4δ::kanr, esc1δ::hphr MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, lrs4δ::kanr, esc1δ::hphr MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, nup84δ::kanr, esc1δ::hphr MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, nup84δ::kanr, esc1δ::hphr MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, nup84δ::kanr, lrs4δ::hphr MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, nup84δ::kanr, lrs4δ::hphr KMY666 MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, heh1δ::kanr KMY682 MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, heh1 ::KanR KMY624 MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, nur1δ::kanr KMY653 MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, nur1δ::kanr KMY838 MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, cik1δ::kanr KMY882 MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, cik1δ::kanr KMY1747 MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, cik1δ::kanr, lrs4 ::HphR S8
9 KMY1770 MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, cik1δ::kanr, lrs4 ::HphR KMY1744 MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, kar3δ::hphr KMY1764 MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, kar3δ::hphr KMY1745 KMY1767 MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, lrs4δ::kanr, kar3 ::HphR MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, lrs4δ::kanr, kar3 ::HphR KMY1617 MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, cin8δ::hphr KMY1629 MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, cin8δ::hphr KMY2247 MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, bub2δ::natr KMY2272 MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, bub2δ::natr KMY780 MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, rad52δ::kanr KMY821 MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, rad52δ::kanr KMY1596 KMY1613 MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, lrs4δ::kanr, rad52δ::hphr MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, lrs4δ::kanr, rad52δ::hphr KMY1225 MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, dnl4δ::kanr KMY1240 MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, dnl4δ::kanr KMY1650 KMY1666 MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, lrs4δ::kanr, dnl4δ::hphr MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, lrs4δ::kanr, dnl4δ::hphr KMY856 MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, ku70δ::kanr KMY1083 MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, ku70δ::kanr KMY2148 MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, pol32δ::hphr KMY2160 MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, pol32δ::hphr KMY2151 MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, dpb3δ::hphr KMY2156 MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, dpb3δ::hphr KMY875 MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, sae2δ::hphr KMY1130 MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, sae2δ::hphr KMY862 MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, sir4δ::hphr KMY1095 MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, sir4δ::hphr KMY852 MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, sir3δ::kanr KMY1089 MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, sir3δ::kanr KMY1148 MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, sir3δ::kanr rad52δ::hphr KMY1178 MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, sir3δ::kanr, S9
10 rad52δ::hphr KMY576 1 MATa, ura3-δ0, leu2δ1, his3δ200, lys2δ202, ykl201c::csura3cs KMY643 MATa, ura3-δ0, leu2δ1, his3δ200, lys2δ202, ykl201c::csura3cs KMY783 MATa, ura3-δ0, leu2δ1, his3δ200, lys2δ202, ykl201c::csura3cs, rad52δ::kanr KMY824 MATa, ura3-δ0, leu2δ1, his3δ200, lys2δ202, ykl201c::csura3cs, rad52δ::kanr KMY859 MATa, ura3-δ0, leu2δ1, his3δ200, lys2δ202, ykl201c::csura3cs, ku70δ::kanr KMY1086 MATa, ura3-δ0, leu2δ1, his3δ200, lys2δ202, ykl201c::csura3cs, ku70δ::kanr KMY1119 MATa, ura3-δ0, leu2δ1, his3δ200, lys2δ202, ykl201c::csura3cs, dnl4δ::kanr KMY1136 MATa, ura3-δ0, leu2δ1, his3δ200, lys2δ202, ykl201c::csura3cs, dnl4δ::kanr KMY779 MATa, ura3-δ0, leu2δ1, his3δ200, lys2δ202, ykl201c::csura3cs, nup84δ::kanr KMY846 MATa, ura3-δ0, leu2δ1, his3δ200, lys2δ202, ykl201c::csura3cs, nup84δ::kanr KMY733 MATa, ura3-δ0, leu2δ1, his3δ200, lys2δ202, ykl201c::csura3cs, esc1δ::hphr KMY760 MATa, ura3-δ0, leu2δ1, his3δ200, lys2δ202, ykl201c::csura3cs, esc1δ::hphr KMY691 MATa, ura3-δ0, leu2δ1, his3δ200, lys2δ202, ykl201c::csura3cs, lrs4δ::kanr KMY714 MATa, ura3-δ0, leu2δ1, his3δ200, lys2δ202, ykl201c::csura3cs, lrs4δ::kanr KMY694 MATa, ura3-δ0, leu2δ1, his3δ200, lys2δ202, ykl201c::csura3cs, csm1δ::kanr KMY718 MATa, ura3-δ0, leu2δ1, his3δ200, lys2δ202, ykl201c::csura3cs, csm1δ::kanr KMY697 MATa, ura3-δ0, leu2δ1, his3δ200, lys2δ202, ykl201c::csura3cs, heh1δ::kanr KMY722 MATa, ura3-δ0, leu2δ1, his3δ200, lys2δ202, ykl201c::csura3cs, heh1δ::kanr KMY699 MATa, ura3-δ0, leu2δ1, his3δ200, lys2δ202, ykl201c::csura3cs, nur1δ::kanr KMY726 MATa, ura3-δ0, leu2δ1, his3δ200, lys2δ202, ykl201c::csura3cs, nur1δ::kanr KMY1595 MATa, ura3-δ0, leu2δ1, his3δ200, lys2δ202, ykl201c::csura3cs, cik1δ::hphr KMY1626 MATa, ura3-δ0, leu2δ1, his3δ200, lys2δ202, ykl201c::csura3cs, cik1δ::hphr KMY865 MATa, ura3-δ0, leu2δ1, his3δ200, lys2δ202, ykl201c::csura3cs, sir4δ::hphr KMY1098 MATa, ura3-δ0, leu2δ1, his3δ200, lys2δ202, ykl201c::csura3cs, sir4δ::hphr KMY854 MATa, ura3-δ0, leu2δ1, his3δ200, lys2δ202, ykl201c::csura3cs, sir3δ::kanr KMY1092 MATa, ura3-δ0, leu2δ1, his3δ200, lys2δ202, ykl201c::csura3cs, sir3δ::kanr KMY1156 MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, slx8δ::hphr KMY1184 MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, slx8δ::hphr KMY1160 MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, slx5δ::hphr KMY1190 MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, slx5δ::hphr KMY1159 MATa, ura3-δ0, leu2δ1, his3δ200, lys2δ202, ykl201c::csura3cs, slx8δ::hphr KMY1187 MATa, ura3-δ0, leu2δ1, his3δ200, lys2δ202, ykl201c::csura3cs, slx8δ::hphr KMY1163 MATa, ura3-δ0, leu2δ1, his3δ200, lys2δ202, ykl201c::csura3cs, slx5δ::hphr KMY1193 MATa, ura3-δ0, leu2δ1, his3δ200, lys2δ202, ykl201c::csura3cs, slx5δ::hphr KMY1171 S10
11 KMY1205 KMY1166 KMY1199 KMY1169 KMY1202 KMY1593 KMY1607 KMY1594 KMY1647 KMY578 1 KMY1683 KMY1684 KMY1780 KMY2011 KMY580 1 KMY2189 KanR KanR MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, Lrs4-Myc13- KanR MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, Lrs4-Myc13- KanR MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, Sir3-Myc13- KanR MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, Sir3-Myc13- KanR KanR, lrs4δ::hphr KanR, lrs4δ::hphr KanR, cik1δ::hphr KanR, cik1δ::hphr MATa, ura3δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3cs::tet0112::natmx, LEU2::tetR-GFP MATa, ura3δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3cs::tet0112::natmx, LEU2::tetR-GFP, Nup49-GFP-KanR MATa, ura3δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3cs::tet0112::natmx, LEU2::tetR-GFP, Nup49-GFP-KanR, lrs4δ::hphr MATa, ura3δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3cs::tet0112::natmx, LEU2::tetR-GFP, Nup49-GFP-KanR, nur1δ::hphr MATa, ura3δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3cs::tet0112::natmx, LEU2::tetR-GFP, Nup49-GFP-KanR, cik1δ::hphr MATa, ura3δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3cs::tet0112::natmx, LEU2::tetR-GFP, nup145cδ::kanmx4 MATa, ura3δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3cs::tet0112::natmx, LEU2::tetR-GFP, nup145cδ::kanmx4, Nup49-GFP-His3 KMY2190 MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, Cik1-Myc-NatR KMY2263 MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, Cik1-Myc-NatR KMY2193 KMY2266 MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, Lrs4Δ::KanR, Cik1-Myc-NatR MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, Lrs4Δ::KanR, Cik1-Myc-NatR KMY219 Heh1-TAP-TRPkl, Lrs4-Myc13-KanR KMY1910 Heh1-TAP-TRPkl, Lrs4-Myc13-KanR KMY2058 MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, Nup84-Myc- NatR, Lrs4-TAP-KanR S11
12 KMY2074 MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, Nup84-Myc- NatR, Lrs4-TAP-KanR KMY2250 MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, rad9δ::natr KMY2275 MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, rad9δ::natr KMY2253 MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, swr1δ::natr KMY2278 MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, swr1δ::natr KMY2256 KMY2281 MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, cik1δ::kanr, swr1δ::natr MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, cik1δ::kanr, swr1δ::natr KMY2012 MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, tub3δ::hphr KMY2027 MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, tub3δ::hphr KMY2018 KMY2033 KMY2258 KMY2289 KMY2261 KMY2292 KMY1738* KMY1861* KMY1733* KMY1849* KMY2076* KMY2078* KMY2009* KMY2067* KMY2050* MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, cik1δ::kanr, tub3δ::hphr MATa, ura3-δ851, leu2δ1, his3δ200, lys2δ202, ykl222c::csura3csa, cik1δ::kanr, tub3δ::hphr KanR, swr1δ::natr KanR, swr1δ::natr KanR, cik1δ::hphr, swr1δ::natr KanR, cik1δ::hphr, swr1δ::natr NatR NatR NatR NatR NatR, Nup84-Myc13-KanR NatR, Nup84-Myc13-KanR NatR, Nup84-Myc13-KanR, lrs4δ::hphr NatR, Nup84-Myc13-KanR, lrs4δ::hphr NatR, Nup84-Myc13-KanR, cik1δ::hphr S12
13 KMY2070* KMY2036* KMY2061* KMY2051* KMY2064* KMY2057* KMY2071* KMY1815* KMY1864* KMY1800* KMY1852* KMY1847* KMY1867* KMY1803* KMY1855* KMY1913* KMY1989* KMY1807* KMY1858* DDY NatR, Nup84-Myc13-KanR, cik1δ::hphr NatR, Nup84-Myc13-KanR NatR, Nup84-Myc13-KanR NatR, Nup84-Myc13-KanR, lrs4δ::hphr NatR, Nup84-Myc13-KanR, lrs4δ::hphr NatR, Nup84-Myc13-KanR, cik1δ::hphr NatR, Nup84-Myc13-KanR, cik1δ::hphr NatR, lrs4δ::hphr NatR, lrs4δ::hphr NatR, lrs4δ::hphr NatR, lrs4δ::hphr NatR, cik1δ::hphr NatR, cik1δ::hphr NatR, cik1δ::hphr NatR, cik1δ::hphr NatR, nup84δ::hphr NatR, nup84δ::hphr NatR, nup84δ::hphr NatR, nup84δ::hphr JRL346 Mata::HOcsDEL::hisG ura3 DEL851 trp1del63 sup53del::leu2del::natmx hmldel::hisg hmrdel::ade3 ade3::gal10::ho can1,1-1446::hocs::hph::del AVT2 ykl215c::leu2::hisg::can1del1-289 DDY3398 JRL346 pol32::kanmx DDY3394 JRL346 cik1::kanmx S13
14 DDY3396 JRL346 kar3::kanmx DDY4009 JRL346 lrs4::kanmx (clone #1) DDY4010 JRL346 lrs4::kanmx (clone #2) DDY4011 JRL346 nup84::kanmx (clone #1) DDY4012 JRL346 nup84::kanmx (clone #2) DDY4013 JRL346 kar3::kanmx prs414 DDY4014 JRL346 kar3::kanmx prs414-kar3 DDY4015 JRL346 kar3::kanmx prs414-kar3-1 DDY4016 JRL346 kar3::kanmx prs414-kar3-898 DDY DDY4008 DDY3094 DDY3095 DDY3093 DDY3096 DDY355 8 DDY3091 DDY3089 DDY3090 DDY3092 DDY DDY3430 DDY3431 DDY DDY3099 YMV45 ho hml::ade1 mata::hisg hmr::ade1 leu2::leu2(asp718-sali)-ura3-pbr332- MATa ade3::gal::ho ade1 lys5 ura3-52 trp1::hisg YMV45 ho hml::ade1 mata::hisg hmr::ade1 leu2::leu2(asp718-sali)-ura3-pbr332- MATa ade3::gal::ho ade1 lys5 ura3-52 trp1::hisg rad52::kanmx YMV45 ho hml::ade1 mata::hisg hmr::ade1 leu2::leu2(asp718-sali)-ura3-pbr332- MATa ade3::gal::ho ade1 lys5 ura3-52 trp1::hisg cik1::kanmx YMV45 ho hml::ade1 mata::hisg hmr::ade1 leu2::leu2(asp718-sali)-ura3-pbr332- MATa ade3::gal::ho ade1 lys5 ura3-52 trp1::hisg kar3::kanmx YMV45 ho hml::ade1 mata::hisg hmr::ade1 leu2::leu2(asp718-sali)-ura3-pbr332- MATa ade3::gal::ho ade1 lys5 ura3-52 trp1::hisg nup84::kanmx YMV45 ho hml::ade1 mata::hisg hmr::ade1 leu2::leu2(asp718-sali)-ura3-pbr332- MATa ade3::gal::ho ade1 lys5 ura3-52 trp1::hisg htz1::kanmx YMV2 MATD::hisG hml::ade1 hmr::ade1 his4::ura3::trp1-leu2 (Xho1-toAsp718)- pbr322-his4 ade1 lys5 ura3-52 trp1::hisg leu2::hocs ade3::gal::ho YMV2 MATD::hisG hml::ade1 hmr::ade1 his4::ura3::trp1-leu2 (Xho1-toAsp718)- pbr322-his4 ade1 lys5 ura3-52 trp1::hisg leu2::hocs ade3::gal::ho rad52::natmx YMV2 MATD::hisG hml::ade1 hmr::ade1 his4::ura3::trp1-leu2 (Xho1-toAsp718)- pbr322-his4 ade1 lys5 ura3-52 trp1::hisg leu2::hocs ade3::gal::ho cik1::natmx YMV2 MATD::hisG hml::ade1 hmr::ade1 his4::ura3::trp1-leu2 (Xho1-toAsp718)- pbr322-his4 ade1 lys5 ura3-52 trp1::hisg leu2::hocs ade3::gal::ho kar3::natmx YMV2 MATD::hisG hml::ade1 hmr::ade1 his4::ura3::trp1-leu2 (Xho1-toAsp718)- pbr322-his4 ade1 lys5 ura3-52 trp1::hisg leu2::hocs ade3::gal::ho nup84::kanmx YMV2 MATD::hisG hml::ade1 hmr::ade1 his4::ura3::trp1-leu2 (Xho1-toAsp718)- pbr322-his4 ade1 lys5 ura3-52 trp1::hisg leu2::hocs ade3::gal::ho htz1::kanmx BY4743 Mata/α his3δ1/his3δ1 leu2δ0/leu2δ0 lys2δ0/lys2 met15δ0/met15 ura3δ 0/ura3 est2::kanmx/est2 cik1::natmx BY4743 Mata/α his3δ1/his3δ1 leu2δ0/leu2δ0 lys2δ0/lys2 met15δ0/met15 ura3δ 0/ura3 est2::kanmx/est2 kar3::natmx TGI354 hoδ MATa-inc arg5,6::mata-hph ade3::gal-ho hmrδ::ade hmlδ ::ADE1 ura3-52 TGI354 hoδ MATa-inc arg5,6::mata-hph ade3::gal-ho hmrδ::ade hmlδ ::ADE1 ura3-52 cik1::natmx DDY3100 TGI354 hoδ MATa-inc arg5,6::mata-hph ade3::gal-ho hmrδ::ade hmlδ ::ADE1 S14
15 ura3-52 rad52::natmx DDY4302 SubTEL Nup49-mCherry::HPH prs413-rad52-yfp DDY4303 SubTEL cik1::kanmx Nup49-mCherry::HPH prs413-rad52-yfp DDY4296 JRL346 Nup49-mCherry::URA3 prs415-rad52-yfp DDY4297 JRL346 Nup49-mCherry::URA3 prs415-rad52-yfp cik1::kanmx DDY4298 JRL346 Nup49-mCherry::URA3 prs415-rad52-yfp kar3::kanmx DDY4299 JRL346 Nup49-mCherry::URA3 prs415-rad52-yfp kar3::kanmx prs414-kar3 DDY4300 JRL346 Nup49-mCherry::URA3 prs415-rad52-yfp kar3::kanmx prs414-kar3-1 DDY4301 JRL346 Nup49-mCherry::URA3 prs415-rad52-yfp kar3::kanmx prs414-empty *cmrs-dsb-l = Control sequence, MRS-DSB-L = zip sequence S15
16 Supplementary Table 4. List of primer pairs used for PCR survivor analysis, ChIP analysis, and gemic integration of NPC-targeting DNA zip/control sequences. Location (Application) Across SubTEL DSB (Survivor) Across SubTEL DSB (zip/cont. survivor) Telomere (Generic AC 1-3 sequence) 0.7 kb left of SubTEL DSB (ChIP) 0.7 kb right of SubTEL DSB (ChIP) 0.07 kb to TEL VI R (ChIP) Control locus ACT1 (ChIP) Amplifying product for transformation (zip integration) Amplifying product for transformation (Control integration) 0.6 kb left of SubTEL DSB (zip/cont. ChIP) 1.5 kb away from n-telomeric DSB (ChIP) Size (bp) (NHEJ) (NHEJ) Sequence 1 Sequence 2 CTGAGTCTGCACTAGACAAT (P1) GAAGTTAAGTGCGCAGAAAG (P1*) ATCTTGATCTCAAAAGCACC (P2) ATCTTGATCTCAAAAGCACC (P2) variable ACCACACACCCACCAC (P3) ATCTTGATCTCAAAAGCACC (P2) 200 TTCTCTCAATACCATCCGC AGAAGCTTATTGTCTAAGCCC 350 GGGAGTAAATTAGACTATGC TAGAATACTTTGCATTCCCA 270 CATGACCAGTCCTCATTTCCATC ACGTTTAGCTGAGTTTAACGGTG 153 GCCTTCTACGTTTCCATCCA GGCCAAATCGATTCTCAAAA n/a n/a CCCAAAAAAAAGAGCAACTATCTAG AAAAGTATGTAGAAGTCCTTCTTTC CCGGATCCCCGGGTTAATTAA CCCAAAAAAAAGAGCAACTATCTAG AAAAGTATGTAGAAGTCGATCATT GCCGGATCCCCGGGTTAATTAA TTTGTGAAAGCTATTGGTGGCACC CTCTGTTCCTTTCGGAGAATTCGAG CTCGTTTAAAC TTTGTGAAAGCTATTGGTGGCACC CTCTGTTCCTTTCGGAGAATTCGAG CTCGTTTAAAC 362 TTCTCACATCACATCCGAACA ATGTCCTCGACGGTCAGC 268 (qpcr only) GTATGACTCACCCGGAAACC ATAACGGCAAACAGCAAAGG S16
17 Supplementary References 1 Therizols, P. et al. Telomere tethering at the nuclear periphery is essential for efficient DNA double strand break repair in subtelomeric region. J. Cell Biol. 172, (2006). 2 Lydeard, J. R., Lipkin-Moore, Z., Jain, S., Eapen, V. V. & Haber, J. E. Sgs1 and exo1 redundantly inhibit break-induced replication and de vo telomere addition at broken chromosome ends. PLoS Genet 6, e (2010). 3 Vaze, M. B. et al. Recovery from checkpoint-mediated arrest after repair of a doublestrand break requires Srs2 helicase. Mol Cell 10, (2002). 4 Yeung, M. & Durocher, D. Srs2 enables checkpoint recovery by promoting disassembly of DNA damage foci from chromatin. DNA Repair (Amst) 10, (2011). 5 Ira, G., Malkova, A., Liberi, G., Foiani, M. & Haber, J. E. Srs2 and Sgs1-Top3 suppress crossovers during double-strand break repair in yeast. Cell 115, (2003). S17
Supplemental Table 1. List of the simple sequence repeat (SSR) and single nucleotide polymorphic (SNP) markers used in the genetic cluster analysis.
 Supplemental Table 1. List of the simple sequence repeat (SSR) and single nucleotide polymorphic (SNP) markers used in the genetic cluster analysis. The markers are listed with the associated wheat chromosome.
Supplemental Table 1. List of the simple sequence repeat (SSR) and single nucleotide polymorphic (SNP) markers used in the genetic cluster analysis. The markers are listed with the associated wheat chromosome.
The Impressive Increase in Throughput of the illumina Genome Analyzer, as Seem from an User Perspective
 The Impressive Increase in Throughput of the illumina Genome Analyzer, as Seem from an User Perspective Laurent FARINELLI 31 August 2009 Illumina Seminar 55 Congresso Brasileiro de Genética Águas de Lindóia,
The Impressive Increase in Throughput of the illumina Genome Analyzer, as Seem from an User Perspective Laurent FARINELLI 31 August 2009 Illumina Seminar 55 Congresso Brasileiro de Genética Águas de Lindóia,
Journal of Avian Biology
 Journal of Avian Biology JAV-00814 Alvarez, S., salter, J. F., McCormack, J. E. and Milá, B. 2015. Speciation in mountain refugia: phylogeography and demographic history of the pine and blackcapped complex.
Journal of Avian Biology JAV-00814 Alvarez, S., salter, J. F., McCormack, J. E. and Milá, B. 2015. Speciation in mountain refugia: phylogeography and demographic history of the pine and blackcapped complex.
Rapid detection and evolutionary analysis of Legionella pneumophila serogroup 1 ST47
 Rapid detection and evolutionary analysis of Legionella pneumophila serogroup 1 ST47 M. Mentasti, P. Cassier, S. David, C. Ginevra, L. Gomez-Valero, A. Underwood, B. Afshar, J. Etienne, Julian Parkhill,
Rapid detection and evolutionary analysis of Legionella pneumophila serogroup 1 ST47 M. Mentasti, P. Cassier, S. David, C. Ginevra, L. Gomez-Valero, A. Underwood, B. Afshar, J. Etienne, Julian Parkhill,
Bi190 Mating Type Interconversion
 Ir Herskowitz Bi190 Mting Type Interconversion 2011 Sternberg, Cltech Scchromyces cerevisie (budding yest) Life Cycle mting / zygote sporultion -C -N Ascus with 4 spores germintion +C +N hploid cells specific
Ir Herskowitz Bi190 Mting Type Interconversion 2011 Sternberg, Cltech Scchromyces cerevisie (budding yest) Life Cycle mting / zygote sporultion -C -N Ascus with 4 spores germintion +C +N hploid cells specific
Bi190 Mating Type Interconversion
 Ir Herskowitz Bi190 Mting Type Interconversion 2007 Sternberg, Cltech Scchromyces cerevisie (budding yest) Life Cycle mting / zygote sporultion -C -N germintion +C +N Ascus with 4 spores hploid cells specific
Ir Herskowitz Bi190 Mting Type Interconversion 2007 Sternberg, Cltech Scchromyces cerevisie (budding yest) Life Cycle mting / zygote sporultion -C -N germintion +C +N Ascus with 4 spores hploid cells specific
Confirmation Protocol for E. coli O157:H7
 Introduction Confirmation Protocol for E. coli O157:H7 The following protocol is used by Hygiena to recover E. coli O157:H7 from beef samples that were enriched according to the BAX System method. The
Introduction Confirmation Protocol for E. coli O157:H7 The following protocol is used by Hygiena to recover E. coli O157:H7 from beef samples that were enriched according to the BAX System method. The
Comparison of Gelman and Millipore Membrane Filters for Enumerating Fecal Coliform Bacteria
 APPLIED MICROBIOLOGY, Sept. 1973, p. 332-336 Copyright 0 1973 American Society for Microbiology Vol. 26, No. 3 Printed in U.S.A. Comparison of Gelman and Millipore Membrane Filters for Enumerating Fecal
APPLIED MICROBIOLOGY, Sept. 1973, p. 332-336 Copyright 0 1973 American Society for Microbiology Vol. 26, No. 3 Printed in U.S.A. Comparison of Gelman and Millipore Membrane Filters for Enumerating Fecal
Comprehensive analysis of SET domain gene family in foxtail millet identifies the putative role of SiSET14 in abiotic stress tolerance
 Comprehensive analysis of SET domain gene family in foxtail millet identifies the putative role of SiSET14 in abiotic stress tolerance Chandra Bhan Yadav, Mehanathan Muthamilarasan, Anand Dangi, Shweta
Comprehensive analysis of SET domain gene family in foxtail millet identifies the putative role of SiSET14 in abiotic stress tolerance Chandra Bhan Yadav, Mehanathan Muthamilarasan, Anand Dangi, Shweta
Journal of Avian Biology
 Journal of Avian Biology JAV-01885 Illera, J. C., Rando, J. C., Rodriguez-Exposito, E., Hernández, M., Claramunt, S. and Martín, A. 2018. Acoustic, genetic, and morphological analysis of the Canarian common
Journal of Avian Biology JAV-01885 Illera, J. C., Rando, J. C., Rodriguez-Exposito, E., Hernández, M., Claramunt, S. and Martín, A. 2018. Acoustic, genetic, and morphological analysis of the Canarian common
Pr oject Summar y. Impact of ground beef packaging systems and temperature abuse on the safety of ground beef
 Pr oject Summar y Impact of ground beef packaging systems and temperature abuse on the safety of ground beef Principal Investigators: J Chance Brooks, Mindy M. Brashears, Mark F. Miller, and Adam Tittor
Pr oject Summar y Impact of ground beef packaging systems and temperature abuse on the safety of ground beef Principal Investigators: J Chance Brooks, Mindy M. Brashears, Mark F. Miller, and Adam Tittor
Using molecular markers for germplasm identification in Bosnia and Herzegovina
 University of Banja Luka Genetic Resources Institute Using molecular markers for germplasm identification in Bosnia and Herzegovina MSc Mirela Kajkut Prof. Dr Gordana Đurić Budapest, 3-5 March 2014 Genetic
University of Banja Luka Genetic Resources Institute Using molecular markers for germplasm identification in Bosnia and Herzegovina MSc Mirela Kajkut Prof. Dr Gordana Đurić Budapest, 3-5 March 2014 Genetic
TIME LIMITS AND MAINTENANCE CHECKS
 TIME LIMITS AND MAINTENANCE CHECKS 1. GENERAL This chapter provides the recommended intervals for the overhaul and replacement of components, and the scheduled and unscheduled maintenance for the airplane.
TIME LIMITS AND MAINTENANCE CHECKS 1. GENERAL This chapter provides the recommended intervals for the overhaul and replacement of components, and the scheduled and unscheduled maintenance for the airplane.
AVERY DENNISON Traffic Sign Fabricator
 STANDARD TRAFFIC SIGN SIZES: 45cm x 45cm 60cm x 60cm 75cm x 75cm 90cm x 90cm 75cm x 75cm 90cm x 90cm 60cm x 60cm 75cm x 75cm 90cm x 90cm 120cm x 120cm 30cm x 30cm 40cm x 40cm 45cm x 45cm 60cm x 60cm 75cm
STANDARD TRAFFIC SIGN SIZES: 45cm x 45cm 60cm x 60cm 75cm x 75cm 90cm x 90cm 75cm x 75cm 90cm x 90cm 60cm x 60cm 75cm x 75cm 90cm x 90cm 120cm x 120cm 30cm x 30cm 40cm x 40cm 45cm x 45cm 60cm x 60cm 75cm
SEG Houston 2009 International Exposition and Annual Meeting
 Constrained propeller ship noise removal and its application to OC data Manhong Guo*, Jun Cai, Jim Specht, in Wang TGS-Nopec Geophysical Company, 500 CityWest lvd. Suite 000, Houston, TX 7704, US Summary
Constrained propeller ship noise removal and its application to OC data Manhong Guo*, Jun Cai, Jim Specht, in Wang TGS-Nopec Geophysical Company, 500 CityWest lvd. Suite 000, Houston, TX 7704, US Summary
Larval fish dispersal in a coral-reef seascape
 VOLUME: 1 ARTICLE NUMBER: 0148 In the format provided by the authors and unedited. Larval fish dispersal in a coral-reef seascape Glenn R. Almany 1, Serge Planes 1, Simon R. Thorrold 2 *, Michael L. Berumen
VOLUME: 1 ARTICLE NUMBER: 0148 In the format provided by the authors and unedited. Larval fish dispersal in a coral-reef seascape Glenn R. Almany 1, Serge Planes 1, Simon R. Thorrold 2 *, Michael L. Berumen
Pr oject Summar y. Survey of the prevalence of Escherichia coli O157:H7 on the surface of subprimal cuts of beef during winter months (Phase I)
 Pr oject Summar y Survey of the prevalence of Escherichia coli O157:H7 on the surface of subprimal cuts of beef during winter months (Phase I) Principal Investigators: J. E. (Ken) Kennedy ABC Research
Pr oject Summar y Survey of the prevalence of Escherichia coli O157:H7 on the surface of subprimal cuts of beef during winter months (Phase I) Principal Investigators: J. E. (Ken) Kennedy ABC Research
Molecular characterization of Italian Soil-borne cereal mosaic virus isolates
 11 Parasitica, 2005, 61(1):11-16 Molecular characterization of Italian Soil-borne cereal mosaic virus isolates C. RATTI 1, A. PISI 1, V. VALLEGA 2 & C. RUBIES AUTONELL 1 1 Department of Agroenvironmental
11 Parasitica, 2005, 61(1):11-16 Molecular characterization of Italian Soil-borne cereal mosaic virus isolates C. RATTI 1, A. PISI 1, V. VALLEGA 2 & C. RUBIES AUTONELL 1 1 Department of Agroenvironmental
Tissue samples, voucher specimens and sequence accession numbers
 Electronic Supplementary Material S1 Internal sequencing primers for partial ND4+tRNA His,Ser,Leu sequence Three internal oligonucleotide primers were designed to sequence overlapping portions of a PCR
Electronic Supplementary Material S1 Internal sequencing primers for partial ND4+tRNA His,Ser,Leu sequence Three internal oligonucleotide primers were designed to sequence overlapping portions of a PCR
Microdeletion FISH Probes
 Microdeletion FISH Probes Microdeletion FISH Probes Oxford Gene Technology offers a comprehensive range of Microdeletion FISH probes from the Cytocell Aquarius portfolio. All Aquarius probes are directly
Microdeletion FISH Probes Microdeletion FISH Probes Oxford Gene Technology offers a comprehensive range of Microdeletion FISH probes from the Cytocell Aquarius portfolio. All Aquarius probes are directly
NOTCH1 mutations are associated with high CD49d expression in Chronic Lymphocytic Leukemia: link between the NOTCH1 and the NF-kB pathways
 NOTCH1 mutations are associated with high CD49d expression in Chronic Lymphocytic Leukemia: link between the NOTCH1 and the NF-kB pathways Supplementary information: Supplementary Materials and Methods;
NOTCH1 mutations are associated with high CD49d expression in Chronic Lymphocytic Leukemia: link between the NOTCH1 and the NF-kB pathways Supplementary information: Supplementary Materials and Methods;
Group constant generation for PARCS using Helios and Serpent and comparison to Serpent 3D model
 6th International Serpent User Group Meeting Politecnico di Milano, Milan, Italy September 26 th -30 th, 2016 Group constant generation for PARCS using Helios and Serpent and comparison to Serpent 3D model
6th International Serpent User Group Meeting Politecnico di Milano, Milan, Italy September 26 th -30 th, 2016 Group constant generation for PARCS using Helios and Serpent and comparison to Serpent 3D model
Operator s Manual. Wedgemaster 606N 808N 808SG ENGLISH
 ENGLISH Wedgemaster 606N 808N 808SG Thank you for purchasing this Vollrath Food Processing Equipment. Before operating the equipment, read and familiarize yourself with the following operating and safety
ENGLISH Wedgemaster 606N 808N 808SG Thank you for purchasing this Vollrath Food Processing Equipment. Before operating the equipment, read and familiarize yourself with the following operating and safety
Assembling & Fitting Instruction August 2015
 Assembling & Fitting Instruction August 2015 Wave Curtain Heading System Silent Gliss 6010, 6020, 6100, 6103, 6290, 6380, 6465 Silent Gliss 3840, 6120 Silent Gliss 5090, 5200, 5400 Silent Gliss 5100, 5600
Assembling & Fitting Instruction August 2015 Wave Curtain Heading System Silent Gliss 6010, 6020, 6100, 6103, 6290, 6380, 6465 Silent Gliss 3840, 6120 Silent Gliss 5090, 5200, 5400 Silent Gliss 5100, 5600
Origin and genetic variation of tree of heaven in Eastern Austria, an area of early introduction
 Institute of Silviculture University of Natural Resources and Life Sciences (BOKU), Vienna Origin and genetic variation of tree of heaven in Eastern Austria, an area of early introduction Vienna, 13.9.2018
Institute of Silviculture University of Natural Resources and Life Sciences (BOKU), Vienna Origin and genetic variation of tree of heaven in Eastern Austria, an area of early introduction Vienna, 13.9.2018
Paragonimus mexicanus Miyazaki e Ishii, 1968
 Morphological and molecular study of three populations of Miyazaki e Ishii, 1968 (Digenea: Paragonimidae) in Mexico Jorge López Caballero 1, Virginia León Règagnon 1, Luis García Prieto 1, David Osorio
Morphological and molecular study of three populations of Miyazaki e Ishii, 1968 (Digenea: Paragonimidae) in Mexico Jorge López Caballero 1, Virginia León Règagnon 1, Luis García Prieto 1, David Osorio
Interpretation Guide
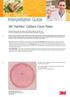 3M Petrifilm Interpretation Guide 3M Petrifilm Coliform Count Plates This guide familiarizes you with results on 3M Petrifilm Coliform Count Plates (CC). For further information, please contact the 3M
3M Petrifilm Interpretation Guide 3M Petrifilm Coliform Count Plates This guide familiarizes you with results on 3M Petrifilm Coliform Count Plates (CC). For further information, please contact the 3M
The Effectiveness of JetBlue if Allowed to Manage More of its Resources
 McNair Scholars Research Journal Volume 2 Article 4 2015 The Effectiveness of JetBlue if Allowed to Manage More of its Resources Jerre F. Johnson Embry Riddle Aeronautical University, johnsff9@my.erau.edu
McNair Scholars Research Journal Volume 2 Article 4 2015 The Effectiveness of JetBlue if Allowed to Manage More of its Resources Jerre F. Johnson Embry Riddle Aeronautical University, johnsff9@my.erau.edu
NORTHERN FEEL.
 NORTHERN FEEL Work Be Creative Get Inspired Nature has always been a source of inspiration and relaxation. Taiga Concept products provide a touch of nature, helping to create a better working environment
NORTHERN FEEL Work Be Creative Get Inspired Nature has always been a source of inspiration and relaxation. Taiga Concept products provide a touch of nature, helping to create a better working environment
Coverage of Mangrove Ecosystem along Three Coastal Zones of Puerto Rico using IKONOS Sensor
 Coverage of Mangrove Ecosystem along Three Coastal Zones of Puerto Rico using IKONOS Sensor Jennifer Toledo Rivera Geology Department, University of Puerto Rico, Mayagüez Campus P.O. Box 9017 Mayagüez,
Coverage of Mangrove Ecosystem along Three Coastal Zones of Puerto Rico using IKONOS Sensor Jennifer Toledo Rivera Geology Department, University of Puerto Rico, Mayagüez Campus P.O. Box 9017 Mayagüez,
Analysis of en-route vertical flight efficiency
 Analysis of en-route vertical flight efficiency Technical report on the analysis of en-route vertical flight efficiency Edition Number: 00-04 Edition Date: 19/01/2017 Status: Submitted for consultation
Analysis of en-route vertical flight efficiency Technical report on the analysis of en-route vertical flight efficiency Edition Number: 00-04 Edition Date: 19/01/2017 Status: Submitted for consultation
Sizing up Australia s eastern Grey Nurse Shark population
 Image: David Harasti A new estimate of adult population size for Australia s eastern Grey Nurse Shark drew on widespread genetic sampling and forensic exploration of family trees. Grey Nurse Sharks are
Image: David Harasti A new estimate of adult population size for Australia s eastern Grey Nurse Shark drew on widespread genetic sampling and forensic exploration of family trees. Grey Nurse Sharks are
Temperature affects the silicate morphology in a diatom
 Supplementary Information: Temperature affects the silicate morphology in a diatom N. Javaheri 1, R. Dries 1&2, A. Burson 3, L.J. Stal 3&4, P.M.A. Sloot 1,5,6, J.A. Kaandorp 1* 1- Computational Science,
Supplementary Information: Temperature affects the silicate morphology in a diatom N. Javaheri 1, R. Dries 1&2, A. Burson 3, L.J. Stal 3&4, P.M.A. Sloot 1,5,6, J.A. Kaandorp 1* 1- Computational Science,
Petrifilm. Interpretation Guide. Coliform Count Plate. Brand
 Petrifilm Brand Interpretation Guide The 3M Petrifilm is a sample-ready culture medium system that contains modified Violet Red Bile nutrients, a cold-water-soluble gelling agent and a tetrazolium indicator
Petrifilm Brand Interpretation Guide The 3M Petrifilm is a sample-ready culture medium system that contains modified Violet Red Bile nutrients, a cold-water-soluble gelling agent and a tetrazolium indicator
This document is a preview generated by EVS
 INTERNATIONAL STANDARD ISO 5912 Fourth edition 2011-10-01 Camping tents Tentes de camping Reference number ISO 5912:2011(E) ISO 2011 ISO 2011 COPYRIGHT PROTECTED DOCUMENT All rights reserved. Unless otherwise
INTERNATIONAL STANDARD ISO 5912 Fourth edition 2011-10-01 Camping tents Tentes de camping Reference number ISO 5912:2011(E) ISO 2011 ISO 2011 COPYRIGHT PROTECTED DOCUMENT All rights reserved. Unless otherwise
Interpretation Guide
 3M Petrifilm Interpretation Guide 3M Petrifilm Coliform Count Plates This guide familiarizes you with results on 3M Petrifilm Coliform Count Plates (CC). For further information, please contact the 3M
3M Petrifilm Interpretation Guide 3M Petrifilm Coliform Count Plates This guide familiarizes you with results on 3M Petrifilm Coliform Count Plates (CC). For further information, please contact the 3M
Flexible Pavement Design
 Flexible Pavement Design FAARFIELD 1.3 Workshop Starting Screen No Job Files Created Click on New Job Presented to: VI ALACPA Airport Pavements Seminar & IV FAA Workshop By: David R. Brill, P.E., Ph.D.
Flexible Pavement Design FAARFIELD 1.3 Workshop Starting Screen No Job Files Created Click on New Job Presented to: VI ALACPA Airport Pavements Seminar & IV FAA Workshop By: David R. Brill, P.E., Ph.D.
Tool: Overbooking Ratio Step by Step
 Tool: Overbooking Ratio Step by Step Use this guide to find the overbooking ratio for your hotel and to create an overbooking policy. 1. Calculate the overbooking ratio Collect the following data: ADR
Tool: Overbooking Ratio Step by Step Use this guide to find the overbooking ratio for your hotel and to create an overbooking policy. 1. Calculate the overbooking ratio Collect the following data: ADR
California Leafy Greens Research Board Final Report April 1, 2008 to March 31, 2009
 California Leafy Greens Research Board Final Report April 1, 28 to March 31, 29 I. Abstract Project Title: Survival of attenuated Escherichia coli O157:H7 ATCC 7728 in fieldinoculated lettuce. Project
California Leafy Greens Research Board Final Report April 1, 28 to March 31, 29 I. Abstract Project Title: Survival of attenuated Escherichia coli O157:H7 ATCC 7728 in fieldinoculated lettuce. Project
Molecular characterization of the Andean blackberry, Rubus glaucus, using SSR markers
 Molecular characterization of the Andean blackberry, Rubus glaucus, using SSR markers M. Marulanda, A.M. López and M. Uribe Laboratorio de Biotecnología Vegetal, Facultad de Ciencias Ambientales, Universidad
Molecular characterization of the Andean blackberry, Rubus glaucus, using SSR markers M. Marulanda, A.M. López and M. Uribe Laboratorio de Biotecnología Vegetal, Facultad de Ciencias Ambientales, Universidad
Takahiro Masuda, Shosuke Iwamoto, Ryohei Yoshinaga, Hidetoshi Tozaki-Saitoh, Akira
 upplementary nformati Trancripti factor drive 2X4 reactive microglia gating neuropathic pain Takahiro Mauda, houke wamoto, yohei Yohinaga, Hidetohi Tozaki-aitoh, Akira Nihiyama, Tak W. Mak, Tomohiko Tamura,
upplementary nformati Trancripti factor drive 2X4 reactive microglia gating neuropathic pain Takahiro Mauda, houke wamoto, yohei Yohinaga, Hidetohi Tozaki-aitoh, Akira Nihiyama, Tak W. Mak, Tomohiko Tamura,
Photopoint Monitoring in the Adirondack Alpine Zone
 Photopoint Monitoring in the Adirondack Alpine Zone Julia Goren (PI) and Seth Jones Adirondack High Peaks Summit Steward Program Adirondack Mountain Club summit@adk.org PO Box 867, Lake Placid, NY 12946
Photopoint Monitoring in the Adirondack Alpine Zone Julia Goren (PI) and Seth Jones Adirondack High Peaks Summit Steward Program Adirondack Mountain Club summit@adk.org PO Box 867, Lake Placid, NY 12946
NordVal International / NMKL c/o Norwegian Veterinary Institute PB 750 Sentrum, 0106 Oslo, Norway
 Issued for: 3M TM Petrifilm TM / Coliform Count Plate NordVal No: 014 First approval date: 5 May 2003 Renewal date: 1 June 2017 Valid until: 1 June 2019 3M TM Petrifilm TM / Coliform Count Plate Manufactured
Issued for: 3M TM Petrifilm TM / Coliform Count Plate NordVal No: 014 First approval date: 5 May 2003 Renewal date: 1 June 2017 Valid until: 1 June 2019 3M TM Petrifilm TM / Coliform Count Plate Manufactured
TSA Pre (PreCheck) Standard Screening. Dwarfism Awareness Month
 Dwarfism Awareness Month TSA Pre (PreCheck) Travelers eligible for TSA Pre (PreCheck): o Proceed to the TSA Pre (PreCheck) line; o Present your boarding pass and government-issued ID to the TSA travel
Dwarfism Awareness Month TSA Pre (PreCheck) Travelers eligible for TSA Pre (PreCheck): o Proceed to the TSA Pre (PreCheck) line; o Present your boarding pass and government-issued ID to the TSA travel
Protecting Consumers. Improving lab efficiency. 3M Petrifilm Plates and Reader
 Protecting Consumers. Improving lab efficiency. 3M Petrifilm Plates and Reader Simply brilliant. This red dot changed microbiology. Imagine what it can do for your lab. In today s environment of heightened
Protecting Consumers. Improving lab efficiency. 3M Petrifilm Plates and Reader Simply brilliant. This red dot changed microbiology. Imagine what it can do for your lab. In today s environment of heightened
Investigation of the effect of antibiotics on bacterial growth. Introduction. Apparatus. Diagram of Apparatus
 Investigation of the effect of antibiotics on bacterial growth Introduction Antimicrobials are agents that are able to kill bacteria or halt their growth. They are widely used in medicine to treat bacterial
Investigation of the effect of antibiotics on bacterial growth Introduction Antimicrobials are agents that are able to kill bacteria or halt their growth. They are widely used in medicine to treat bacterial
Virulence Profiles of Vibrio vulnificus in German Coastal Waters, a Comparison of North Sea and Baltic Sea Isolates
 Supplementary Information Virulence Profiles of Vibrio vulnificus in German Coastal Waters, a Comparison of North Sea and Baltic Sea Isolates Table S1. Sampling sites, classification and bathing water
Supplementary Information Virulence Profiles of Vibrio vulnificus in German Coastal Waters, a Comparison of North Sea and Baltic Sea Isolates Table S1. Sampling sites, classification and bathing water
Aircraft No. 2 N917AU, T-17; P3-A; Serial Number
 Aircraft No. 2 N917AU, T-17; P3-A; Serial Number 150510 Aircraft Data: Manufacture Date: 2/18/1963 Entered airtanker operations: 9/16/2008 FAA Registration Card - See Attachment 2-1. Issued to: Aero Union
Aircraft No. 2 N917AU, T-17; P3-A; Serial Number 150510 Aircraft Data: Manufacture Date: 2/18/1963 Entered airtanker operations: 9/16/2008 FAA Registration Card - See Attachment 2-1. Issued to: Aero Union
The Economic Impact of Tourism Brighton & Hove Prepared by: Tourism South East Research Unit 40 Chamberlayne Road Eastleigh Hampshire SO50 5JH
 The Economic Impact of Tourism Brighton & Hove 2014 Prepared by: Tourism South East Research Unit 40 Chamberlayne Road Eastleigh Hampshire SO50 5JH CONTENTS 1. Summary of Results 1 1.1 Introduction 1 1.2
The Economic Impact of Tourism Brighton & Hove 2014 Prepared by: Tourism South East Research Unit 40 Chamberlayne Road Eastleigh Hampshire SO50 5JH CONTENTS 1. Summary of Results 1 1.1 Introduction 1 1.2
TrueView Features. Amanda J. Minnich and Dr. Abdullah Mueen University of New Mexico {aminnich, March 9, 2015
 TrueView Features Amanda J. Minnich and Dr. Abdullah Mueen University of New Mexico {aminnich, mueen}@cs.unm.edu March 9, 2015 This is a description of the features used in our outlier detection algorithms
TrueView Features Amanda J. Minnich and Dr. Abdullah Mueen University of New Mexico {aminnich, mueen}@cs.unm.edu March 9, 2015 This is a description of the features used in our outlier detection algorithms
Registry Publication 17
 Preparation Requirements for Certificate of Airworthiness (C of A) Renewal Survey The following are the preparation requirements to enable the Operator (normally the person identified on Form 31 as the
Preparation Requirements for Certificate of Airworthiness (C of A) Renewal Survey The following are the preparation requirements to enable the Operator (normally the person identified on Form 31 as the
Zipping the Three Nephews. Klaus Mayer MIPS, Helmholtz Center Munich
 Zipping the Three Nephews Klaus Mayer MIPS, Helmholtz Center Munich Brachypodium Genome Synteny Rice Sorghum Maize r1 r2 r3 r4 r5 r6 r7 r8 r9 r10 r11 r12 s1 s2 s3 s4 s5 s6 s7 s8 s9 s10 m1 m2 m3 m4 m5 m6
Zipping the Three Nephews Klaus Mayer MIPS, Helmholtz Center Munich Brachypodium Genome Synteny Rice Sorghum Maize r1 r2 r3 r4 r5 r6 r7 r8 r9 r10 r11 r12 s1 s2 s3 s4 s5 s6 s7 s8 s9 s10 m1 m2 m3 m4 m5 m6
TEACHER PAGE Trial Version
 TEACHER PAGE Trial Version * After completion of the lesson, please take a moment to fill out the feedback form on our web site (https://www.cresis.ku.edu/education/k-12/online-data-portal)* Lesson Title:
TEACHER PAGE Trial Version * After completion of the lesson, please take a moment to fill out the feedback form on our web site (https://www.cresis.ku.edu/education/k-12/online-data-portal)* Lesson Title:
Pr oject Summar y. Colonization characteristics of bovine recto-anal junction tissues by Escherichia coli O157:H7
 Pr oject Summar y Colonization characteristics of bovine recto-anal junction tissues by Escherichia coli O157:H7 Principal Investigators: James L Bono, Terrance M. Arthur, and Tommy L. Wheeler U.S. Department
Pr oject Summar y Colonization characteristics of bovine recto-anal junction tissues by Escherichia coli O157:H7 Principal Investigators: James L Bono, Terrance M. Arthur, and Tommy L. Wheeler U.S. Department
Measuring Performance of an Automated and Miniaturized LANCE Ultra camp Assay for the G i -coupled 5-HT 1A Receptor a Comparative Study
 application Note Time-Resolved Fluorescence Resonance Energy Transfer Authors Mireille Caron Julie Blouin Nancy Gauthier Philippe Roby Lucille Beaudet Jaime Padrόs PerkinElmer, Inc. Montreal, QC, Canada
application Note Time-Resolved Fluorescence Resonance Energy Transfer Authors Mireille Caron Julie Blouin Nancy Gauthier Philippe Roby Lucille Beaudet Jaime Padrόs PerkinElmer, Inc. Montreal, QC, Canada
Operating Instructions
 Operating Instructions High Throughput Dialysis MODEL HTD96b www.htdialysis.com HTD96b : Operating Instructions Components Universal Base Unit Teflon block assembly: 9 Teflon bars, labeled sequentially
Operating Instructions High Throughput Dialysis MODEL HTD96b www.htdialysis.com HTD96b : Operating Instructions Components Universal Base Unit Teflon block assembly: 9 Teflon bars, labeled sequentially
[Docket No. FAA ; Directorate Identifier 2010-NM-107-AD; Amendment ; AD ]
![[Docket No. FAA ; Directorate Identifier 2010-NM-107-AD; Amendment ; AD ] [Docket No. FAA ; Directorate Identifier 2010-NM-107-AD; Amendment ; AD ]](/thumbs/95/125067740.jpg) [Federal Register Volume 76, Number 37 (Thursday, February 24, 2011)] [Rules and Regulations] [Pages 10215-10216] From the Federal Register Online via the Government Printing Office [www.gpo.gov] [FR Doc
[Federal Register Volume 76, Number 37 (Thursday, February 24, 2011)] [Rules and Regulations] [Pages 10215-10216] From the Federal Register Online via the Government Printing Office [www.gpo.gov] [FR Doc
Sikorsky S 61. Overhaul & Repair Capabilities. USAerospacetn.com. U.S. Aerospace Corp. FAA Certified Part 145 Repair Station License USCR533K
 Sikorsky S 61 Overhaul & Repair Capabilities FAA Certified Part 145 Repair Station License USCR533K EASA Approved Certificate No. EASA.145.6330 USAerospacetn.com Effective: May 1, 2014 U.S. Aerospace Corp.
Sikorsky S 61 Overhaul & Repair Capabilities FAA Certified Part 145 Repair Station License USCR533K EASA Approved Certificate No. EASA.145.6330 USAerospacetn.com Effective: May 1, 2014 U.S. Aerospace Corp.
Supplementary Materials Figures
 Supplementary Materials Figures!"#$%&'(%)**$*$! +,-%!."$/01-0,2! +,-%!!"#$.,30-04$ 5)*3$ +$
Supplementary Materials Figures!"#$%&'(%)**$*$! +,-%!."$/01-0,2! +,-%!!"#$.,30-04$ 5)*3$ +$
CMX521: THE FIRST NUCLEOSIDE IN CLINICAL DEVELOPMENT FOR NOROVIRUS. Randall Lanier, PhD
 CMX521: THE FIRST NUCLEOSIDE IN CLINICAL DEVELOPMENT FOR NOROVIRUS Randall Lanier, PhD Norovirus Infection Is Prevalent and Costly Worldwide: ~700 million cases of norovirus each year (~20 million in U.S.)
CMX521: THE FIRST NUCLEOSIDE IN CLINICAL DEVELOPMENT FOR NOROVIRUS Randall Lanier, PhD Norovirus Infection Is Prevalent and Costly Worldwide: ~700 million cases of norovirus each year (~20 million in U.S.)
A proven system. Paris, France. Aeronest, Lake Neuchatel, Switzerland. Paris, France
 A proven system Paris, France Aeronest, Lake Neuchatel, Switzerland Paris, France November 2002 Safety At Aerophile safety is a continuing obsession. Everything is designed to guarantee the highest possible
A proven system Paris, France Aeronest, Lake Neuchatel, Switzerland Paris, France November 2002 Safety At Aerophile safety is a continuing obsession. Everything is designed to guarantee the highest possible
Have Descents Really Become More Efficient? Presented by: Dan Howell and Rob Dean Date: 6/29/2017
 Have Descents Really Become More Efficient? Presented by: Dan Howell and Rob Dean Date: 6/29/2017 Outline Introduction Airport Initiative Categories Methodology Results Comparison with NextGen Performance
Have Descents Really Become More Efficient? Presented by: Dan Howell and Rob Dean Date: 6/29/2017 Outline Introduction Airport Initiative Categories Methodology Results Comparison with NextGen Performance
Supplemental Information
 Neuron, Volume 88 Supplemental Information Time-Resolved Imaging Reveals Heterogeneous Landscapes of Nanomolar Ca 2+ in Neurons and Astroglia Kaiyu Zheng, Lucie Bard, James P. Reynolds, Claire King, Thomas
Neuron, Volume 88 Supplemental Information Time-Resolved Imaging Reveals Heterogeneous Landscapes of Nanomolar Ca 2+ in Neurons and Astroglia Kaiyu Zheng, Lucie Bard, James P. Reynolds, Claire King, Thomas
COMPARATIVE STUDY ON GROWTH AND FINANCIAL PERFORMANCE OF JET AIRWAYS, INDIGO AIRLINES & SPICEJET AIRLINES COMPANIES IN INDIA
 Volume 2, Issue 2, November 2017, ISBR Management Journal ISSN(Online)- 2456-9062 COMPARATIVE STUDY ON GROWTH AND FINANCIAL PERFORMANCE OF JET AIRWAYS, INDIGO AIRLINES & SPICEJET AIRLINES COMPANIES IN
Volume 2, Issue 2, November 2017, ISBR Management Journal ISSN(Online)- 2456-9062 COMPARATIVE STUDY ON GROWTH AND FINANCIAL PERFORMANCE OF JET AIRWAYS, INDIGO AIRLINES & SPICEJET AIRLINES COMPANIES IN
Supplemental Table 1. TCRβ repertoire of naïve D b PA TRBV29 + CD8 + T cells in B6 mice. CDR3β M1 M2 M3 M4 M5 M6 SWGGEQ SWGERL 2 - -
 Supplemental Table 1. TCRβ repertoire of naïve D b PA + 224 TRBV29 + CD8 + T cells in B6 mice. CDR3β M1 M2 M3 M4 M5 M6 SWGGEQ 2-1 - 1 - SWGERL 2 - - - 1 - SPDRGAL 2 - - - - - SLGDEQ 1 - - - 1 1 AAGREQ
Supplemental Table 1. TCRβ repertoire of naïve D b PA + 224 TRBV29 + CD8 + T cells in B6 mice. CDR3β M1 M2 M3 M4 M5 M6 SWGGEQ 2-1 - 1 - SWGERL 2 - - - 1 - SPDRGAL 2 - - - - - SLGDEQ 1 - - - 1 1 AAGREQ
Abstract. Introduction
 COMPARISON OF EFFICIENCY OF SLOT ALLOCATION BY CONGESTION PRICING AND RATION BY SCHEDULE Saba Neyshaboury,Vivek Kumar, Lance Sherry, Karla Hoffman Center for Air Transportation Systems Research (CATSR)
COMPARISON OF EFFICIENCY OF SLOT ALLOCATION BY CONGESTION PRICING AND RATION BY SCHEDULE Saba Neyshaboury,Vivek Kumar, Lance Sherry, Karla Hoffman Center for Air Transportation Systems Research (CATSR)
Bacterial Quality of Crystalline Rock and Glacial Aquifers in New England
 Bacterial Quality of Crystalline Rock and Glacial Aquifers in New England By Sarah Flanagan and Charles Culbertson, U.S. Geological Survey 2012 New Hampshire Water and Watershed Conference Plymouth State
Bacterial Quality of Crystalline Rock and Glacial Aquifers in New England By Sarah Flanagan and Charles Culbertson, U.S. Geological Survey 2012 New Hampshire Water and Watershed Conference Plymouth State
Biol (Fig 6.13 Begon et al) Logistic growth in wildebeest population
 Biol 303 1 Interspecific Competition Outline Intraspecific competition = density dependence Intraspecific and interspecific competition Limiting resources Interference vs exploitation Effects on population
Biol 303 1 Interspecific Competition Outline Intraspecific competition = density dependence Intraspecific and interspecific competition Limiting resources Interference vs exploitation Effects on population
Get A Grip! Student Activity 1D Introduction: Materials: (per group) LESSON 2
 Get A Grip! Student Activity 1D Introduction: Think for a minute about all of the sports and daily activities that depend on a strong grip! Baseball, bowling, golf, gymnastics, football, hockey, mountain
Get A Grip! Student Activity 1D Introduction: Think for a minute about all of the sports and daily activities that depend on a strong grip! Baseball, bowling, golf, gymnastics, football, hockey, mountain
Comparison on the Ways of Airworthiness Management of Civil Aircraft Design Organization
 Available online at www.sciencedirect.com Procedia Engineering Procedia Engineering 00 (2011) 17 000 000 (2011) 388 395 Procedia Engineering www.elsevier.com/locate/procedia The 2nd International Symposium
Available online at www.sciencedirect.com Procedia Engineering Procedia Engineering 00 (2011) 17 000 000 (2011) 388 395 Procedia Engineering www.elsevier.com/locate/procedia The 2nd International Symposium
A.3 List of Effective Pages
 A.3 List of Effective Pages 1/10 Contents 0 Change History and Overview... 3 1 A General... 4 2 B Management and Administration... 5 3 C Airport Premises... 6 4 D Aviation Data Publication Requirements...
A.3 List of Effective Pages 1/10 Contents 0 Change History and Overview... 3 1 A General... 4 2 B Management and Administration... 5 3 C Airport Premises... 6 4 D Aviation Data Publication Requirements...
Construction of Conflict Free Routes for Aircraft in Case of Free Routing with Genetic Algorithms.
 Construction of Conflict Free Routes for Aircraft in Case of Free Routing with Genetic Algorithms. Ingrid Gerdes, German Aerospace Research Establishment, Institute for Flight Guidance, Lilienthalplatz
Construction of Conflict Free Routes for Aircraft in Case of Free Routing with Genetic Algorithms. Ingrid Gerdes, German Aerospace Research Establishment, Institute for Flight Guidance, Lilienthalplatz
China Aeromodelling Design Challenge. Contest Rules China Aeromodelling Design Challenge Page 1 of 14
 China Aeromodelling Design Challenge Contest Rules 2014 Page 1 of 14 LIST OF CONTENTS I VTOL AIR CARGO RACE... 3 1 OBJECTIVES... 3 2 REGISTRATION ELIGIBILITIES... 3 3 AIRCRAFT CONFIGURATIONS... 3 4 SITE
China Aeromodelling Design Challenge Contest Rules 2014 Page 1 of 14 LIST OF CONTENTS I VTOL AIR CARGO RACE... 3 1 OBJECTIVES... 3 2 REGISTRATION ELIGIBILITIES... 3 3 AIRCRAFT CONFIGURATIONS... 3 4 SITE
Aircraft No. 3 N921AU, T-21; P3-A; Serial Number
 Aircraft No. 3 N921AU, T-21; P3-A; Serial Number 151385 Aircraft Data: Manufacture Date: 1964 Entered airtanker operations: 5/21/1992 FAA Registration Card - See Attachment 3-1. Issued to: Aero Union Corporation
Aircraft No. 3 N921AU, T-21; P3-A; Serial Number 151385 Aircraft Data: Manufacture Date: 1964 Entered airtanker operations: 5/21/1992 FAA Registration Card - See Attachment 3-1. Issued to: Aero Union Corporation
3M TM Petrifilm TM. Petrifilm TM 3M TM. 3M TM Petrifilm TM Serie 2000 Rapid Coliform Count Plates - Ref.: / 50 Unit - Ref.
 3M TM Aerobic Count Plates - Ref.: 06400 / 100 Unit - Ref.: 06406 / 1000 Unit 3M TM Enterobacteriaceae Count Plates 3M TM Coliform Count Plates - Ref.: 06420 / 50 Unit - Ref.: 06421 / 1000 Unit - Ref.:
3M TM Aerobic Count Plates - Ref.: 06400 / 100 Unit - Ref.: 06406 / 1000 Unit 3M TM Enterobacteriaceae Count Plates 3M TM Coliform Count Plates - Ref.: 06420 / 50 Unit - Ref.: 06421 / 1000 Unit - Ref.:
ASSEMBLY 39TH SESSION
 International Civil Aviation Organization WORKING PAPER 19/8/16 ASSEMBLY 39TH SESSION TECHNICAL COMMISSION Agenda Item 37: Other issues to be considered by the Technical Commission TO DEFINE THE VALIDATION
International Civil Aviation Organization WORKING PAPER 19/8/16 ASSEMBLY 39TH SESSION TECHNICAL COMMISSION Agenda Item 37: Other issues to be considered by the Technical Commission TO DEFINE THE VALIDATION
Joe Randell President and Chief Executive Officer Jolene Mahody Executive Vice President and Chief Financial Officer
 Joe Randell President and Chief Executive Officer Jolene Mahody Executive Vice President and Chief Financial Officer Nathalie Megann Vice President, Investor Relations and Corporate Affairs December, 2015
Joe Randell President and Chief Executive Officer Jolene Mahody Executive Vice President and Chief Financial Officer Nathalie Megann Vice President, Investor Relations and Corporate Affairs December, 2015
Summi triumphum. & bc. w w w Ó w w & b 2. Qui. w w w Ó. w w. w w. Ó œ. Let us recount with praise the triumph of the highest King, 1.
 Sequence hymn for Ascension ( y Nottker Balulus) Graduale Patavienese 1511 1. Sum Summi triumphum Let us recount ith praise the triumph of the highest King, Henricus Isaac Choralis Constantinus 1555 3
Sequence hymn for Ascension ( y Nottker Balulus) Graduale Patavienese 1511 1. Sum Summi triumphum Let us recount ith praise the triumph of the highest King, Henricus Isaac Choralis Constantinus 1555 3
Wishlist Plug-in USER GUIDE
 support@simicart.com Phone: 084.4.8585.4587 Wishlist Plug-in USER GUIDE Table of Contents 1. INTRODUCTION... 3 2. HOW TO INSTALL... 4 3. HOW TO CONFIGURE... 5 4. HOW TO USE WISHLIST PLUG-IN... 6 Wishlist
support@simicart.com Phone: 084.4.8585.4587 Wishlist Plug-in USER GUIDE Table of Contents 1. INTRODUCTION... 3 2. HOW TO INSTALL... 4 3. HOW TO CONFIGURE... 5 4. HOW TO USE WISHLIST PLUG-IN... 6 Wishlist
Integration of the Airport and the Network DPI/FUM Messages Management Overview
 Integration of the Airport and the Network DPI/FUM Messages Management Overview Hans Koolen NMOC/CFMU Network Operations Services Edition March 2013 Introduction Purpose The purpose of this presentation
Integration of the Airport and the Network DPI/FUM Messages Management Overview Hans Koolen NMOC/CFMU Network Operations Services Edition March 2013 Introduction Purpose The purpose of this presentation
Advisory Circular. 1.1 Purpose Applicability Description of Changes... 2
 Advisory Circular Subject: Issuing Office: Standards Document No.: AC 521-006 File Classification No.: Z 5000-34 Issue No.: 01 RDIMS No.: 5611040-V40 Effective Date: 2012-03-16 1.1 Purpose... 2 1.2 Applicability...
Advisory Circular Subject: Issuing Office: Standards Document No.: AC 521-006 File Classification No.: Z 5000-34 Issue No.: 01 RDIMS No.: 5611040-V40 Effective Date: 2012-03-16 1.1 Purpose... 2 1.2 Applicability...
INTERPRETATION GUIDE AN INTRODUCTION TO USE AND INTERPRETING RESULTS FOR PEEL PLATE CC TESTS. FOR MORE INFORMATION, CONTACT CHARM SCIENCES
 INTERPRETATION GUIDE AN INTRODUCTION TO USE AND INTERPRETING RESULTS FOR PEEL PLATE CC TESTS. FOR MORE INFORMATION, CONTACT CHARM SCIENCES INTRODUCTION Peel Plate CC (Coliform Count) tests diffuse the
INTERPRETATION GUIDE AN INTRODUCTION TO USE AND INTERPRETING RESULTS FOR PEEL PLATE CC TESTS. FOR MORE INFORMATION, CONTACT CHARM SCIENCES INTRODUCTION Peel Plate CC (Coliform Count) tests diffuse the
PRAJWAL KHADGI Department of Industrial and Systems Engineering Northern Illinois University DeKalb, Illinois, USA
 SIMULATION ANALYSIS OF PASSENGER CHECK IN AND BAGGAGE SCREENING AREA AT CHICAGO-ROCKFORD INTERNATIONAL AIRPORT PRAJWAL KHADGI Department of Industrial and Systems Engineering Northern Illinois University
SIMULATION ANALYSIS OF PASSENGER CHECK IN AND BAGGAGE SCREENING AREA AT CHICAGO-ROCKFORD INTERNATIONAL AIRPORT PRAJWAL KHADGI Department of Industrial and Systems Engineering Northern Illinois University
Environmental restrictions and the efficiency of airports - the case of slot restrictions at Dusseldorf Airport -
 Environmental restrictions and the efficiency of airports - the case of slot restrictions at Dusseldorf Airport - 5th Conference INFRADAY GARS TU Berlin 07 October 06 Hansjochen Ehmer, Thorsten Heidelmeier
Environmental restrictions and the efficiency of airports - the case of slot restrictions at Dusseldorf Airport - 5th Conference INFRADAY GARS TU Berlin 07 October 06 Hansjochen Ehmer, Thorsten Heidelmeier
Display of 1 no. internally illuminated advertisement hoarding
 Committee Date: 22/08/2013 Application Number: 2013/04695/PA Accepted: 01/07/2013 Application Type: Advertisement Target Date: 26/08/2013 Ward: Ladywood Summer Row, Birmingham, B3 1JU Display of 1 no.
Committee Date: 22/08/2013 Application Number: 2013/04695/PA Accepted: 01/07/2013 Application Type: Advertisement Target Date: 26/08/2013 Ward: Ladywood Summer Row, Birmingham, B3 1JU Display of 1 no.
Using The Approach Planner
 Using The Approach Planner photo Living With Your Plane For airports and airfields without published procedures (All graphics in this tutorial are for illustration purposes only and not for flying) A Product
Using The Approach Planner photo Living With Your Plane For airports and airfields without published procedures (All graphics in this tutorial are for illustration purposes only and not for flying) A Product
DF-99 ANTARCTIC PENINSULA SEDIMENT TRAP DEPLOYMENT AND WATER CHEMISTRY CRUISE R/V N.B. PALMER (MAR 28, 1999 to APR. 12, 1999) David A.
 DF-99 ANTARCTIC PENINSULA SEDIMENT TRAP DEPLOYMENT AND WATER CHEMISTRY CRUISE R/V N.B. PALMER (MAR 28, 1999 to APR. 12, 1999) by David A. Mucciarone S-072 Cruise Participant: Robert B. Dunbar and David
DF-99 ANTARCTIC PENINSULA SEDIMENT TRAP DEPLOYMENT AND WATER CHEMISTRY CRUISE R/V N.B. PALMER (MAR 28, 1999 to APR. 12, 1999) by David A. Mucciarone S-072 Cruise Participant: Robert B. Dunbar and David
Typical senario s that may be encountered on a cross country and often constitute the crux points of a flight.
 Module 4a Transition: referring to that part of a flight which entails getting from one place to another (in the most efficient way through effective height management) Typical senario s that may be encountered
Module 4a Transition: referring to that part of a flight which entails getting from one place to another (in the most efficient way through effective height management) Typical senario s that may be encountered
Interpretation Guide 3M Petrifilm Rapid Coliform Count Plates
 3M Petrifilm Interpretation Guide 3M Petrifilm Rapid Coliform Count Plates This guide should familiarize you with results on Petrifilm Rapid Coliform Count (RCC) plates as defined by three of the most
3M Petrifilm Interpretation Guide 3M Petrifilm Rapid Coliform Count Plates This guide should familiarize you with results on Petrifilm Rapid Coliform Count (RCC) plates as defined by three of the most
--- BIG LEAGUE BATTING CAGE ---
 --- BIG LEAGUE BATTING CAGE --- BGLC-7500 Call Jaypro Sports Equipment at 1-800-243-0533 during regular business hours for technical support. www.jaypro.com Rev-B Page 1 of 22 IMPORTANT NOTICE: 1. BEFORE
--- BIG LEAGUE BATTING CAGE --- BGLC-7500 Call Jaypro Sports Equipment at 1-800-243-0533 during regular business hours for technical support. www.jaypro.com Rev-B Page 1 of 22 IMPORTANT NOTICE: 1. BEFORE
ACCORD GENERAL SUR RESTRICTED ET LE COMMERCE. Agreement on Trade in Civil Aircraft Original: English/ anglais TECHNICAL SUB-COMMITTEE.
 GENERAL AGREEMENT ON TARIFFS AND TRADE ACCORD GENERAL SUR RESTRICTED AIR/TSC/W/24/Rev.1 LES TARIFS DOUANIERS24 March 1982 Special Distribution ET LE COMMERCE Agreement on Trade in Civil Aircraft Original:
GENERAL AGREEMENT ON TARIFFS AND TRADE ACCORD GENERAL SUR RESTRICTED AIR/TSC/W/24/Rev.1 LES TARIFS DOUANIERS24 March 1982 Special Distribution ET LE COMMERCE Agreement on Trade in Civil Aircraft Original:
Supporting Information
 Supporting Information Rovito et al. 10.1073/pnas.0813051106 SI Text RT-PCR Batrachochytrium dendrobatidis Assay. This assay uses species-specific primers ITS1 3 Chytr and 5.8S Chytr and the probe ChytrMGB2
Supporting Information Rovito et al. 10.1073/pnas.0813051106 SI Text RT-PCR Batrachochytrium dendrobatidis Assay. This assay uses species-specific primers ITS1 3 Chytr and 5.8S Chytr and the probe ChytrMGB2
TOYOTA TUNDRA BEDRUG Preparation
 TOYOTA TUNDRA 2007 - BEDRUG Preparation Part Number(s): PTS12-34070 (5.5 bed) PTS12-34071 (6.5 bed) PTS12-34072 (8.1 bed) NOTE: Part number of this accessory may not be the same as the part number shown.
TOYOTA TUNDRA 2007 - BEDRUG Preparation Part Number(s): PTS12-34070 (5.5 bed) PTS12-34071 (6.5 bed) PTS12-34072 (8.1 bed) NOTE: Part number of this accessory may not be the same as the part number shown.
The Hiroshima bombing: What you need to know about the nuclear attack
 The Hiroshima bombing: What you need to know about the nuclear attack By Los Angeles Times, adapted by Newsela staff on 06.02.16 Word Count 845 In this Sept. 8, 1945, file photo, a correspondent stands
The Hiroshima bombing: What you need to know about the nuclear attack By Los Angeles Times, adapted by Newsela staff on 06.02.16 Word Count 845 In this Sept. 8, 1945, file photo, a correspondent stands
Airplane Value Analysis Alex Philip
 Airplane Value Analysis Alex Philip Istanbul Technical University Air Transportation Management M.Sc. Program Fundamentals of Airline Management Module 7: 14 October 2015 Financial evaluation of projects
Airplane Value Analysis Alex Philip Istanbul Technical University Air Transportation Management M.Sc. Program Fundamentals of Airline Management Module 7: 14 October 2015 Financial evaluation of projects
MAURITIUS CANE INDUSTRY AUTHORITY MAURITIUS SUGARCANE INDUSTRY RESEARCH INSTITUTE
 MAURITIUS CANE INDUSTRY AUTHORITY MAURITIUS SUGARCANE INDUSTRY RESEARCH INSTITUTE Ref A 1/215 13 August 215 SUGAR CANE CROP 215 Status: End July 215 1. CLIMATE 1.1 Rainfall (Tables 1a and 1b, Figure 1)
MAURITIUS CANE INDUSTRY AUTHORITY MAURITIUS SUGARCANE INDUSTRY RESEARCH INSTITUTE Ref A 1/215 13 August 215 SUGAR CANE CROP 215 Status: End July 215 1. CLIMATE 1.1 Rainfall (Tables 1a and 1b, Figure 1)
Supplemental Figure S1. Co-localization of GH3-2 and putative disease resistance QTLs. LOD, logarithm of odds.
 C922 Resistance to M. grisea isolate F1814 LOD: 2.2 C922 Resistance to Xoo strain JL691 LOD: 1.6 RG101 G393 WRKY13 GH3-2 RG101 G393 WRKY13 GH3-2 RM212 C567 RM212 Chr1 0 0.6 1.2 1.8 2.4 3.0 LOD Chr1 0 0.4
C922 Resistance to M. grisea isolate F1814 LOD: 2.2 C922 Resistance to Xoo strain JL691 LOD: 1.6 RG101 G393 WRKY13 GH3-2 RG101 G393 WRKY13 GH3-2 RM212 C567 RM212 Chr1 0 0.6 1.2 1.8 2.4 3.0 LOD Chr1 0 0.4
Northern genetic richness and southern purity, but just one species in the Chelonoidis chilensis complex
 Northern genetic richness and southern purity, but just one species in the Chelonoidis chilensis complex UWE FRITZ, LEANDRO ALCALDE, MARIO VARGAS-RAMÍREZ, ERIC V. GOODE, DAVID URI FABIUS-TUROBLIN & PETER
Northern genetic richness and southern purity, but just one species in the Chelonoidis chilensis complex UWE FRITZ, LEANDRO ALCALDE, MARIO VARGAS-RAMÍREZ, ERIC V. GOODE, DAVID URI FABIUS-TUROBLIN & PETER
10x Corrosion Resistance 5x Abrasion Resistance 2x Chip Resistance 2x Scratch Resistance. 10x 5x 2x 2x. Textured Powder Paint Finish
 On-Site Storage The latest JOBOX innovations for unmatched security, versatility and durability New Textured Powder Paint Finish For Improved Durability* 10x 5x 2x 2x Site-Vault Security System 50% more
On-Site Storage The latest JOBOX innovations for unmatched security, versatility and durability New Textured Powder Paint Finish For Improved Durability* 10x 5x 2x 2x Site-Vault Security System 50% more
SACAA INTERIM REPORT ON AIRLINK GEORGE AIRPORT ACCIDENT
 SACAA INTERIM REPORT ON AIRLINK GEORGE AIRPORT ACCIDENT Midrand - Following an Airlink accident that took place at the George Airport on 07 December 2009, the South African Civil Aviation Authority s Accident
SACAA INTERIM REPORT ON AIRLINK GEORGE AIRPORT ACCIDENT Midrand - Following an Airlink accident that took place at the George Airport on 07 December 2009, the South African Civil Aviation Authority s Accident
